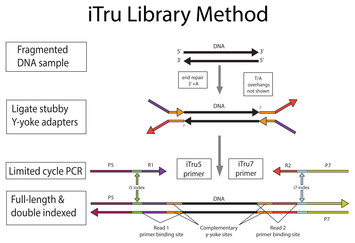
These indexes can generally work with other library preps. This allows samples to be pooled and then individually identified during downstream analysis. Both RNA and DNA preparation kits include adapters containing unique index sequences that are ligated to sample fragments at the beginning of the library construction process. The same TruSeq kit can be used to prepare samples for single-read, paired-end, and multiplexed sequencing on all Illumina sequencing instruments. The universal, methylated adapter design incorporates an index sequence at the initial ligation step. TruSeq library prep uses adapter-embedded indexes to enable high throughput processing and application flexibility. These index adapters cannot be used with other library preps. The PCR step adds index adapter sequences on both ends of the DNA, which enables dual-indexed sequencing of pooled libraries on Illumina sequencing platforms. A limited-cycle PCR step uses the adapters to amplify the insert DNA. (3) Cluster regeneration, add Read 2 sequencing primer to sequence the second read (top strand as template, sequence cDNA):ĥ'- AATGATACGGCGACCACCGAGATCTACACGCCTGTCCGCGG AAGCAGTGGTATCAACGCAGAGTACGT NNNNNNNNNNNN NNNNNNNN(dT)XXX.Nextera library prep uses an engineered transposome to tagment genomic DNA, which is a process that fragments DNA and then tags the DNA with adapter sequences in a single step. Illumina Adapter Sequences Document 1000000002694v06 5 February 2018 Introduction This document lists the index adapter sequences for Illumina library prep kits. (2) Add i7 index sequencing primer to sequence the i7 index (bottom strand as template):ĥ'- CTGTCTCTTATACACATCT CCGAGCCCACGAGAC-> |-5'- TTTTTTTT AAGCAGTGGTATCAACGCAGAGTACGT (dT)XXX.XXX CTGTCTCTTATACACATCTĪAAAAAAA TTCGTCACCATAGTTGCGTCTCATGCA (pA)XXX.XXXXXXXXXXXX GACAGAGAATATGTGTAGA GGCTCGGGTGCTCTG -5'ģ'- TTACTATGCCGCTGGTGGCTCTAGATGTGCGGACAGGCGCC TTCGTCACCATAGTTGCGTCTCATGCA NNNNNNNNNNNN NNNNNNNN(pA)XXX.XXX GACAGAGAATATGTGTAGA GGCTCGGGTGCTCTGNNNNNNNN TAGAGCATACGGCAGAAGACGAAC -5' TTCGTCACCATAGTTGCGTCTCACTTACCCXXXXX.XXXXX(dT) TGCA TGAGACGCAACTATGGTGACGAATTTTTTTT -5'-| Step-by-step library generation (1) mRNA capture using Beads-oligo-dT in the droplets:ĥ'- AAGCAGTGGTATCAACGCAGAGTGAATGGGXXXXX.XXXXX(pA) ACGT ACTCTGCGTTGATACCACTGCTTAAAAAAAA Nextera XT Index Kit v2 adapter Trim bases from sequences2, 3 Trim bases. I7 index sequencing primer: 5'- CTGTCTCTTATACACATCT CCGAGCCCACGAGAC -3' Read 2 sequencing primer: 5'- GTCTCGTGGGCTCGG AGATGTGTATAAGAGACAG -3' Nextera NexteraXT Large/complexgenomes Smallgenomes,amplicons,plasmids Humangenomes PCRAmplicons(>300bp) Nonhumanmammaliangenomes(egmouse, rat,bovine). Read 1 sequencing primer (seqB): 5'- GCCTGTCCGCGG AAGCAGTGGTATCAACGCAGAGTAC -3' Read 1 sequencing primer (seqA): 5'- GCCTGTCCGCGG AAGCAGTGGTATCAACGCAGAGTACGT -3' Library PCR primer 2 (this is basically Nextera N7xx): 5'- CAAGCAGAAGACGGCATACGAGAT GTCTCGTGGGCTCGG -3' 
Library PCR primer 1: 5'- AATGATACGGCGACCACCGAGATCTACACGCCTGTCCGCGG AAGCAGTGGTATCAACGCAGAGTAC -3' 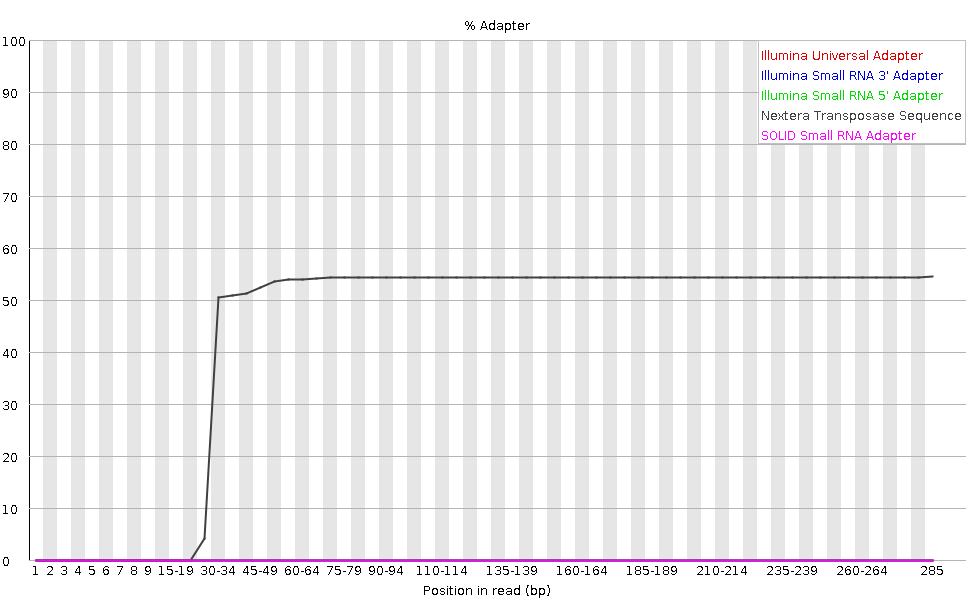
Nextera technology employs in vitro transposition to simultaneously fragment and tag DNA in a single-tube reaction, and prepare sequencer-ready libraries in under 2 hours. Bias: The transposase used in the Nextera system carries mutations and is used under conditions that result in near-random integration. Illumina P7 adapter: 5'- CAAGCAGAAGACGGCATACGAGAT -3' The NexteraTM DNA Sample Prep Kit is designed to prepare genomic DNA libraries compatible with the Illumina Genome Analyzer I and II and Hi- SeqTM 2000 sequencers. EPICENTRE Nextera DNA Sample Prep Kit (Illumina-compatible) page 4 6. Illumina P5 adapter: 5'- AATGATACGGCGACCACCGAGATCTACAC -3' Nextera N7xx primer entry point (s7): 5'- GTCTCGTGGGCTCGG -3' Nextera N/S5xx primer entry point (s5): 5'- TCGTCGGCAGCGTC -3' Nextera Tn5 binding site (19-bp Mosaic End (ME)): 5'- AGATGTGTATAAGAGACAG -3' Trim Galore will try to auto-detect whether the Illumina universal, Nextera transposase or Illumina small RNA adapter sequence. Template Switching Oligo (TSO): 5'- AAGCAGTGGTATCAACGCAGAGTGAATrGrGrG -3' Seq-Well used exact the same oligo design with Drop-seq with Beads-oligo-dT-seqB, which was published in Nature Methods 14, 395–398 (2017).īeads-oligo-dT-seqA: |-5'- TTTTTTT AAGCAGTGGTATCAACGCAGAGTACGT TTTTTTTTTTTTTTTTTTTTTTTTTTTTTT -3'īeads-oligo-dT-seqB: |-5'- TTTTTTT AAGCAGTGGTATCAACGCAGAGTAC TTTTTTTTTTTTTTTTTTTTTTTTTTTTTT -3' Here, Beads-oligo-dT-seqA was used as demonstration. This will produce reads in the desired RF orientation although the alignable length of one read will again be reduced after adapter Data Processing of. In the original pulication in Cell 161, 1202-1214 (2015), there are two batches of beads, with only two base pairs difference. Double-stranded DNA adapter containing long-range PCR primer binding site, sequencing primer binding site, and end marker sequence.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
